Parsing examples#
Some examples on parsing to and from supported formats. More info about all parsing methods can be found in the manual section.
Declare paths#
First, let's do all the necessary imports and generate the paths that we'll use for file import and export.
[1]:
%matplotlib inline
from pathlib import Path
import pygaps.parsing as pgp
# Get directory paths
base_path = Path.cwd() / 'data' / 'parsing'
# Find files
aif_file_paths = list((base_path / 'aif').rglob('*.aif'))
json_file_paths = list((base_path / 'json').rglob('*.json'))
xl_file_paths = list((base_path / 'excel').rglob('*.xls'))
csv_file_paths = list((base_path / 'csv').rglob('*.csv'))
Manufacturer import#
Many report files from various adsorption device manufacturers can be imported directly using pyGAPS. Here are some examples.
[2]:
cfld = base_path / "commercial"
dvssms = pgp.isotherm_from_commercial(cfld / "smsdvs" / "13X water 30c.xlsx", 'smsdvs', 'xlsx')
micromeritics = pgp.isotherm_from_commercial(cfld / "mic" / "Sample_C.xls", 'mic', 'xl')
belsorp_dat = pgp.isotherm_from_commercial(cfld / "bel" / "DUT-13-CH4-190K.DAT", 'bel', 'dat')
threeP_xl = pgp.isotherm_from_commercial(cfld / "3p" / "MOF_N2_77K.xlsx", '3p', 'xl')
quantachrome = pgp.isotherm_from_commercial(cfld / "qnt" / "DUT-6_N2_77K (Raw Analysis Data).txt", 'qnt', 'txt-raw')
Specified adsorbate is not in internal list (or name cannot be resolved to an existing one). Thermodynamic backend disabled for this gas/vapour.
No data collected for pressure_saturation in file c:\Users\pauli\git\pyGAPS\docs\examples\data\parsing\commercial\mic\Sample_C.xls.
AIF Parsing#
AIF Import#
Adsorption information files are fully supported in pyGAPS, both for import and exports. Isotherms can be imported from an .aif as:
[3]:
# Import all
isotherms = [pgp.isotherm_from_aif(path) for path in aif_file_paths]
# Display an example file
print(isotherms[1])
Material: DMOF
Adsorbate: ethane
Temperature: 298.15K
Units:
Uptake in: cm3(STP)/g
Pressure in: kPa
Other properties:
user: single gas
date: 2019-08-19T00:00:00
instrument: BEL VC-05
material_mass: 0.817
material_batch: [Zn2(tm-bdc)2(dabco)]
material_mass_unit: g
AIF Export#
Similarly, an isotherm can be exported as an AIF file or a string, depending on whether a path is passed. For this purpose use either the module pygaps.isotherm_to_aif() function or the convenience class function to_aif().
[4]:
# module function
for isotherm, path in zip(isotherms, aif_file_paths):
pgp.isotherm_to_aif(isotherm, path)
# save to file with convenience function
isotherms[0].to_aif('isotherm.aif')
# string
isotherm_string = isotherms[0].to_aif()
JSON Parsing#
JSON Import#
Isotherms can be imported either from a json file or from a json string. The same function is used in both cases.
[5]:
# Import them
isotherms = [pgp.isotherm_from_json(path) for path in json_file_paths]
# Display an example file
print(isotherms[1])
Material: TEST
Adsorbate: n-butane
Temperature: 298.15K
Units:
Uptake in: g/g
Pressure in: bar
Other properties:
iso_type: isotherm
material_batch: TB
JSON Export#
Exporting to JSON can be done to a file or a string, depending on whether a path is passed. For this purpose use either the module pygaps.isotherm_to_json() function or the convenience class function to_json().
[6]:
# module function
for isotherm, path in zip(isotherms, json_file_paths):
pgp.isotherm_to_json(isotherm, path, indent=4)
# save to file with convenience function
isotherms[0].to_json('isotherm.json')
# string
isotherm_string = isotherms[0].to_json()
Excel Parsing#
Excel does not have to be installed on the system in use.
Excel Import#
[7]:
# Import them
isotherms = [pgp.isotherm_from_xl(path) for path in xl_file_paths]
# Display an example file
print(isotherms[1])
Material: MCM-41
Adsorbate: nitrogen
Temperature: 77.0K
Units:
Uptake in: mmol/g
Pressure in: bar
Other properties:
comment: None
date: None
lab: MADIREL
instrument: Triflex
project: None
activation_temperature: 150.0
user: PI
iso_type: Isotherme
material_batch: Test
[8]:
isotherms[1].plot()
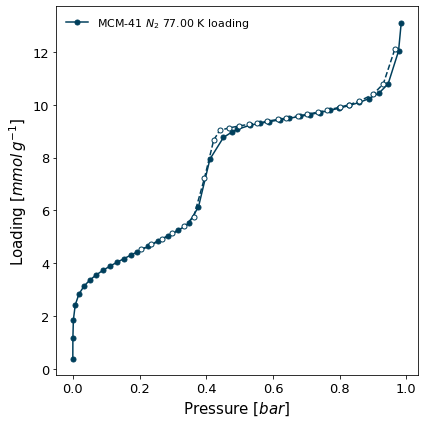
Excel Export#
[9]:
# Export each isotherm in turn
for isotherm, path in zip(isotherms, xl_file_paths):
pgp.isotherm_to_xl(isotherm, path)
# save to file with convenience function
isotherms[0].to_xl('isotherm.xls')
CSV Parsing#
CSV Import#
Like JSON, isotherms can be imported either from a CSV file or from a CSV string. The same function is used in both cases.
[10]:
# Import them
isotherms = [pgp.isotherm_from_csv(path) for path in csv_file_paths]
# Display an example file
print(isotherms[0])
Material: HKUST-1(Cu)
Adsorbate: carbon dioxide
Temperature: 303.0K
Units:
Uptake in: mmol/g
Pressure in: bar
Other properties:
material_batch: Test
iso_type: Calorimetrie
user: ADW
machine: CV
date: 21/05/2010 00:00
activation_temperature: 150.0
lab: MADIREL
CSV Export#
[11]:
# Export each isotherm in turn
for isotherm, path in zip(isotherms, csv_file_paths):
pgp.isotherm_to_csv(isotherm, path)
# save to file with convenience function
isotherms[0].to_csv('isotherm.csv')
# string representation
isotherm_string = isotherms[0].to_csv()